IADR Abstract Archives
Host-microbial interactions of the peri-implant metatranscriptome in health and disease
Objectives: It is estimated that nearly 2 million endosseous dental implants are being placed each year. While implants have proven highly reliable, two microbially driven inflammatory diseases often accompany these restorations: Peri-implant mucositis (affecting 50% of implants in 80% of patients) and peri-implantitis (10% of implants in 20% of patients). Clinical outcomes have been modest with disturbingly high rates of recurrence, and research has been approached using tools of limited scope. The purpose of the present investigation was to characterize host-bacterial interactions in healthy and diseased peri-implant tissues using a dual RNA-Seq approach.
Methods: Systemically healthy subjects with two single implants (one healthy, one diseased) were recruited. Peri-implant mucosal tissue and plaque samples were taken. Host and bacterial total RNA extracted, converted to cDNA and sequenced on the Illumina HiSeq platform. Host transcripts were aligned and quantified against the human genome (GRCh38) using the splicing-aware alignment software kallisto. Bacterial transcripts were matched against HOMD, and MEGAN used to analyze pathways with the KEGG database.
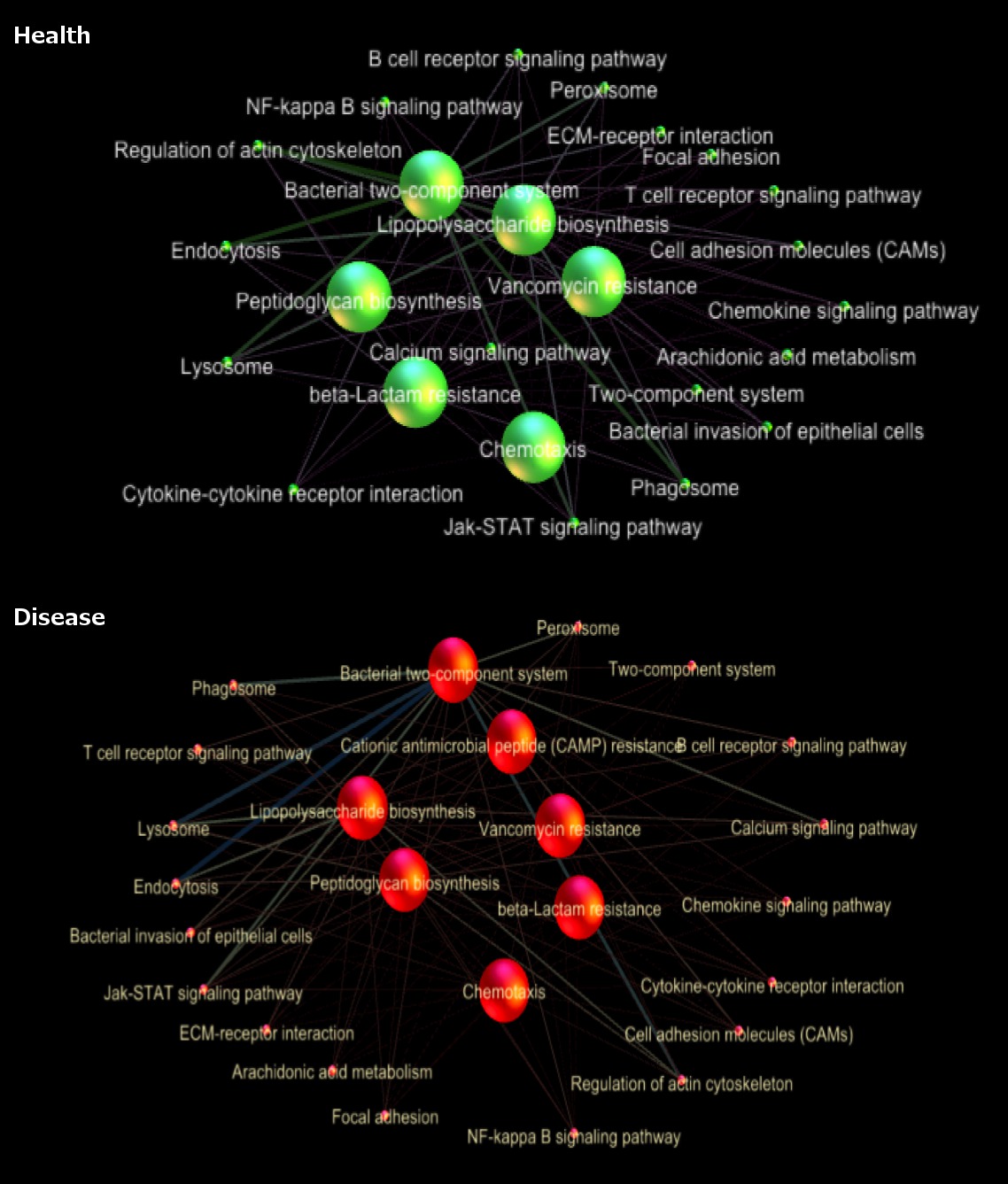
Results: In the host, diseased samples exhibited significantly greater expression of pathways involved in cell-signaling, immune-response, cell-proliferation, and metabolism. In the microbiome, pathways related to membrane-transport, LPS, peptidoglycan biosynthesis, antibiotic-resistance, and chemotaxis were significantly upregulated. Through network analysis of host-bacterial interactions, significant correlations between gene families associated with host immune regulatory signaling pathways, cell-communication, chemokine and cytokine signaling pathways, and bacterial virulence gene families such as LPS biosynthesis, and various antibiotic-resistance functions (vancomycin, beta-lactams, cationic antimicrobial peptide resistances) were revealed in disease.
Conclusions: Here we present the first unbiased open-ended, high throughput -omics approach to study implant disease as it relates to both the host as well as the microbiota. These highly sensitive assays have allowed us to characterize the molecular profile behind these processes and identify potential biomarkers, which will ultimately translate to better clinical outcomes.
Methods: Systemically healthy subjects with two single implants (one healthy, one diseased) were recruited. Peri-implant mucosal tissue and plaque samples were taken. Host and bacterial total RNA extracted, converted to cDNA and sequenced on the Illumina HiSeq platform. Host transcripts were aligned and quantified against the human genome (GRCh38) using the splicing-aware alignment software kallisto. Bacterial transcripts were matched against HOMD, and MEGAN used to analyze pathways with the KEGG database.
Results: In the host, diseased samples exhibited significantly greater expression of pathways involved in cell-signaling, immune-response, cell-proliferation, and metabolism. In the microbiome, pathways related to membrane-transport, LPS, peptidoglycan biosynthesis, antibiotic-resistance, and chemotaxis were significantly upregulated. Through network analysis of host-bacterial interactions, significant correlations between gene families associated with host immune regulatory signaling pathways, cell-communication, chemokine and cytokine signaling pathways, and bacterial virulence gene families such as LPS biosynthesis, and various antibiotic-resistance functions (vancomycin, beta-lactams, cationic antimicrobial peptide resistances) were revealed in disease.
Conclusions: Here we present the first unbiased open-ended, high throughput -omics approach to study implant disease as it relates to both the host as well as the microbiota. These highly sensitive assays have allowed us to characterize the molecular profile behind these processes and identify potential biomarkers, which will ultimately translate to better clinical outcomes.